A new study our lab published in Nature Plants challenges the long-held assumption on how enhancers, key regulatory elements in gene expression, function in plants. It was discovered that unlike their counterparts in humans, enhancers in plants are rarely associated with unstable RNA transcripts.
Enhancers are DNA sequences that can boost the activity of genes, playing a crucial role in various biological processes. In animals, particularly humans, unstable RNA produced from distal bidirectional regions have been widely used as markers for enhancers. However, the new research reveals a distinct pattern in plants.
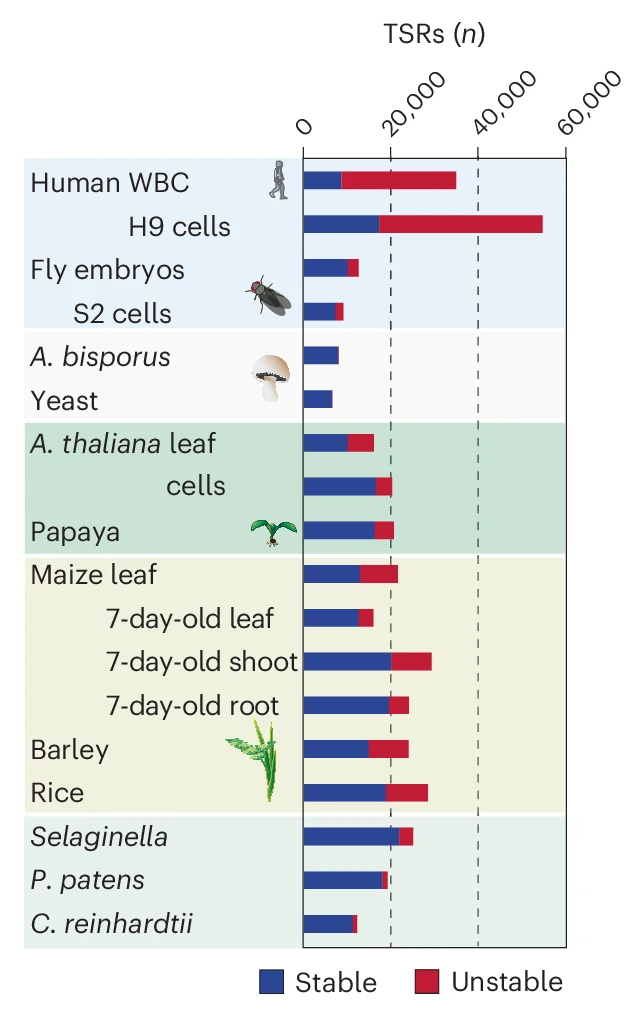
The study analyzed a wide range of plant species and found that unstable RNA transcripts originating from enhancer-like regions are significantly less common compared to humans. Instead, many plant enhancers appear to function more like promoters, initiating stable RNA transcripts.
This groundbreaking discovery has significant implications for our understanding of plant biology and gene regulation. By revealing the unique characteristics of plant enhancers, the research opens up new avenues for exploring gene regulation.