Modeling & Simulation Lab
The major research focus of our lab is to understand the role of membrane lipids in the molecular recognition of therapeutically important integral membrane proteins such as GPCRs, ion channels, and integrins in atomistic detail and to design/discover small-molecule therapeutics using state-of-the-art computational techniques.
Membrane-facilitated ligand access and binding to G Protein-coupled receptors (GPCRs) and Ligand-Gated Ion Channels
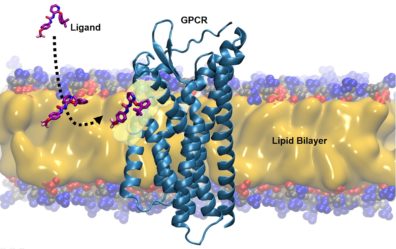
Membrane proteins constitute approximately one-third of all proteins expressed in humans and are involved in numerous fundamental cellular processes. Strikingly, G protein-coupled receptors (GPCRs) and ion channels, together, constitute 31% of all human therapeutic targets through which >50% of the marketed small-molecule drugs exert their pharmacological effects, underlining their significance. Cell membranes have been known to serve as essential structural media for proper localization, structure, and function of these protein targets. However, a growing body of evidence shows that membrane lipids play roles that are even more sophisticated. Remarkably, several crystal structures of GPCRs and ligand-gated ion channels bound to their ligands offer unprecedented details on the existence of allosteric binding sites at lipid-transmembrane helix interfaces that can be accessed only via lipid-mediated paths. In addition, recent biophysical and computational studies (including data from our lab) reveal that lipophilic and amphiphilic ligands gain access to their protein targets by first partitioning into the surrounding membrane and then undergoing two-dimensional lateral diffusion through the membrane. The structural organization and constraints of the bilayer, including lipid composition, appear to play critical roles in directing ligands to a specific depth within the bilayer, and in “pre-organizing” them in an optimal orientation and conformation to facilitate receptor binding. Despite the clear participation of membrane lipids in molecular recognition processes, the mechanistic details of lipid involvement in the binding of drugs to integral membrane proteins remain poorly understood. There is a critical gap in the fundamental knowledge of how interactions among these species affect access, binding kinetics, and affinities of drugs to lipid-exposed transmembrane sites. The long-term goal of our research program is to contribute to this body of knowledge using advanced modeling and simulation techniques.
Outside-in activation of integrins by oxysterols
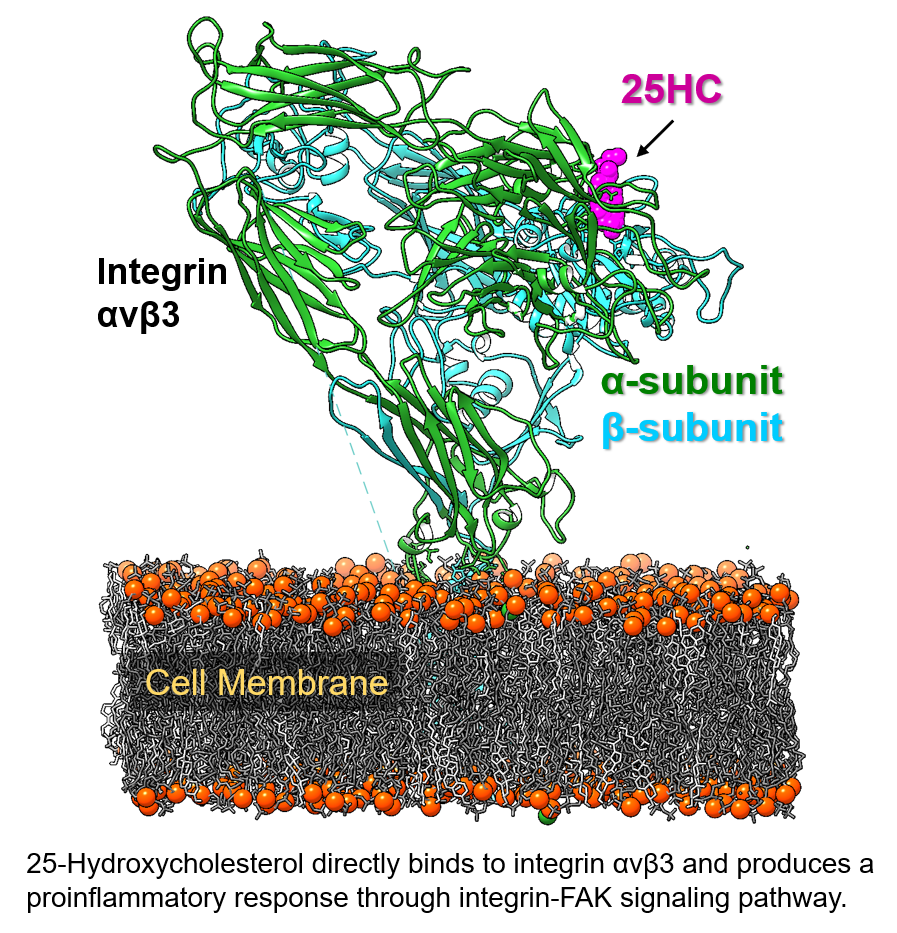
Oxysterols are oxygenated metabolites of cholesterol formed in the human body and are involved in a plethora of physiological and pathological processes such as lipid homeostasis, inflammation, innate and adaptive immunity, cancer, and degenerative brain diseases. Specifically, 25-hydroxycholesterol (25HC) is now established as an important regulator of the immune system and is produced by immune cells in response to viral infection and activation of pattern recognition receptors. Recently, we uncovered a novel cellular mechanism of 25HC-mediated regulation of the proinflammatory response. We showed that 25HC amplifies the activation of immune cells and increases the production of immune mediators such as TNF and IL-6, by directly binding to αvβ3 and α5β1 integrins and activating the integrin-focal adhesion kinase pathway. We also discovered that 25HC binds to integrins at a novel binding site (site 2), distinct from the site where the extracellular matrix (ECM) ligands containing an Arg-Gly-ASP (RGD) motif are known to bind. The binding of 25HC at site 2 produces significant conformational changes in the specificity-determining loop (SDL) of integrins, near the RGD-binding site. The effect of such conformational changes in the SDL on the binding of ECM ligands, as well as the basis of 25HC-mediated allosteric signaling mechanisms underlying integrin activation, are not known. Our central hypothesis is that binding of 25HC to integrins at site 2 triggers conformational changes in the SDL, resulting in an efficient binding of ECM ligands producing a further surge of innate inflammatory response. Our rationale is that small molecule modulators blocking 25HC-integrin interaction would serve as an efficient anti-inflammatory therapeutic strategy to combat various inflammatory diseases. Accordingly, the overall objective of this project is to elucidate the molecular mechanisms of integrin activation by oxysterols and to identify selective small-molecule modulators targeting site 2 of integrins for potential therapeutic applications.
The ongoing research projects are supported by the following grants from the National Institutes of Health.
NIH/NIGMS – R01 GM137022-01 (2020 – 2024)
Molecular biophysics of integrin activation by oxysterols and rational discovery of small-molecule modulators.
Role: PI, in collaboration with Dr. Santanu Bose (Co-I)
NIH/NIGMS – R15 GM131293-01 (2018 – 2021)
Specifics of “Non-Specific Membrane Interactions” of Drugs: An Integrated Approach for Understanding Structure-Membrane Interaction Relationships.
Role: PI
NIH/NIAID – R01 AI083387-07A1 (2018 – 2022)
Molecular and cellular mechanisms regulating innate immunity and inflammation during pattern recognition receptor activation and respiratory virus infection.
Role: Co-I, in collaboration with Dr. Santanu Bose (PI)
Completed Projects
NIH/NINDS – R41 NS107099-01 (2018 – 2019)
In Silico Identification of Novel GHB Receptor Ligands for SSADH Deficiency, a Disorder of GABA Metabolism.
Role: PI, in collaboration with Speragen Inc.

Brian Mun, Charles Mariasoosai, Christopher Szlenk, Senthil Natesan, Adian Talib, Ayobami Diyaolu, and Peter Obi (from left to right)